 
Run the differential equation solver for each patient in par.data #Compile DES function - it's called by lsoda for each individual in the dataset #Function containing differential equations for amount in each compartmentĭA = RateC -K12*A +K21*A -K10*A #Central compartmentĭA = K12*A -K21*A #Peripheral compartment #Do not repeat a time so use "unique" as well #The time sequence must include all "event" times for deSolve so add them here and sort At TIME = 0.16, RATE = 0 (infusion stopped)Ĭstep.doseinf <- approxfun(CTIMEinf, CRATEinf, method = "const") #Define an interpolation function that returns rate when given time - "const" #Creating continuous infusion (for 24 hours) - this use the approxfun function to make a "forcing function" for infusion rate in the differential equationsĬTIMEinf <- c(0,CTinf,24) #100 - make sure the function works long after the 24 hours #There are a number of ways that doses can be coded for the deSolve package - see help for deSolve Make a time sequence and specify the dose information for a system of differential equations #Calculate rate-constants for differential equation solver #Now use multivariate rnorm to turn the covariance matrix into ETA valuesĮTAmat <- mvrnorm(n = n, mu = c(0,0,0,0), OMEGA) #Use this function to turn CORR and SDVAL into a covariance matrix 
#Specify the between subject variability for CL, V1, V2 R24 <- 0 #Correlation coefficient for V1-Q R14 <- 0 #Correlation coefficient for CL-Q R13 <- 0 #Correlation coefficient for CL-V2 QPOP <- 5.7451 #Inter-compartmental clearance L/hĬOV1 <- 0.520245 #Covariate effect of CRCL on clearance (power model)ĬOV2 <- 1.22484 #Covariate effect of gender on CLĬOV3 <-1.51075 #Covariate effect of ALB on V1 Par.data <- seq(from = 1, to = n, by = 1) Create a parameter dataframe with ID and parameter values for each individual Stat3 <- quantile(x, probs=CIhi, names=F) Stat2 <- quantile(x, probs=CIlow, names=F) #Function for calculating median, upper and lower confidence intervals for x #Assign probabilities for 95% confidence intervals #Assign probabilities for 90% confidence intervals #Assign probabilities for 80% confidence intervals #This is called whenever the input changes #Reactive expression to generate a reactive data frame #Define user-input dependent functions for output #TIME range - times where a concentration will be calculated Theme_custom <- theme_set(theme_grey(18)) #Registers the parallel backend with the foreach package (automatically loaded when parallel argument in ddply which calls a function that usesĬlusterEvalQ(cl, list(library(deSolve), library(foreach))) #The foreach package always needs to be included #List packages that are required to be sent to each core for the parallel process #Contents with makePSOCKcluster brackets can be changed to a whole number if you #detectCores() searches for the number of cores that the local machine has #Set up a cluster of cores to run the application overĬl <- makePSOCKcluster(detectCores() - 1) #Setting up cores to run parallel processes, thus increasing speed #Code for functions and variables which are not reactive (not dependent on "input$X") 
PlotOutput("plotCONC", height = 650, width = 750),Īlign = "center") #Brackets closing "mainPanel" 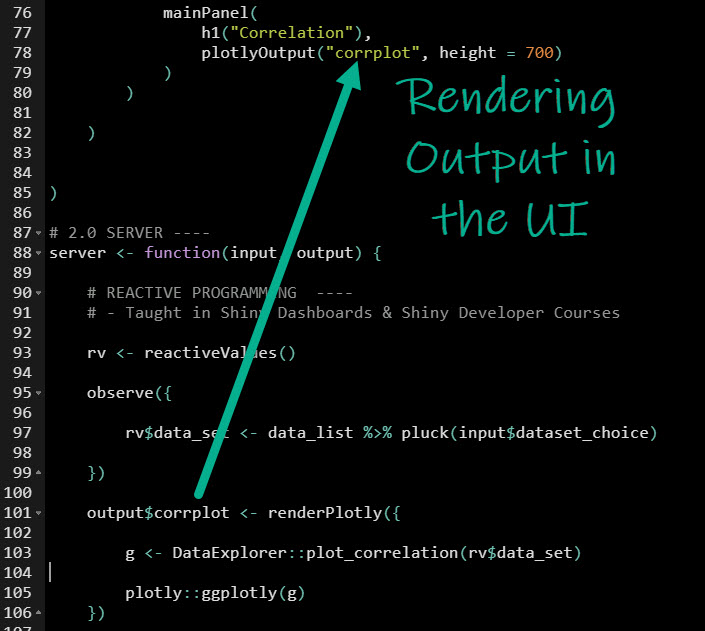
#Plot output for concentration-time profile "95% - 2.5th and 97.5th percentiles" = 3),Īlign = "left"), #Brackets closing "sidebarPanel" "Dose administered as short infusion (mg):",Ĭhoices = list("80% - 10th and 90th percentiles" = 1, Hr(), #Add a break with a horizontal line The UI contains all the parameters that could be altered and used in the model simulation which will show up as a plot. I'm running a differential equation containing model for a drug and something seems to be amiss. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
